RTKfusion.py
Philip Peffer
BACKGROUND:
The vascular endothelial growth factor (VEGF) signaling pathway is critical to the growth and proliferation of cancer cells, as it activates the production of new blood vessels (i.e. angiogenesis) required to bring oxygen to the tumor (Holash et al., 2002). VEGF overexpression is associated with many types of cancer, as well as other diseases, such as wet/neovascular age-related macular degeneration (AMD) (Hicklin & Ellis, 2005; Spilsbury et al., 2000). Because of this, the VEGF pathway has been the subject of much research, and there are currently several VEGF-inhibiting drugs on the market (Papadopoulos et al., 2012; Park et al., 2015).
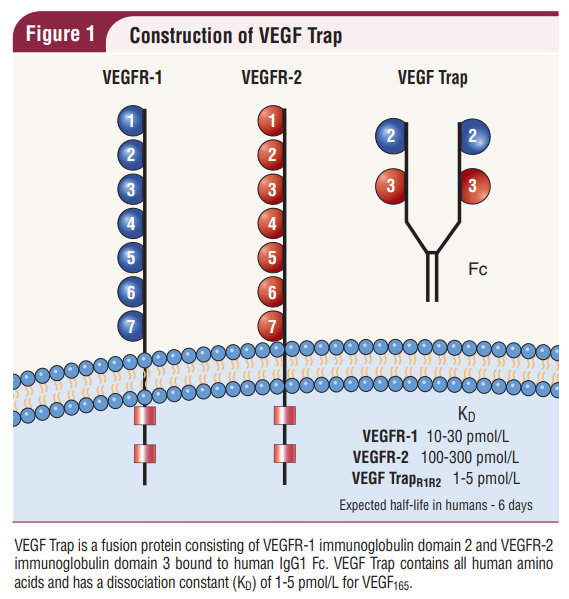
One of these VEGF blockers / decoy receptors, which I work with in my job as a QC biochemist, is VEGF-Trap. VEGF-Trap is a recombinant fusion protein consisting of the immunoglobulin-like domain 2 (Ig2) of VEGF receptor 1 (VEGFR1) , VEGFR2 Ig3, and the constant Fc portion of IgG1 (Ig2&3) fused together (Holash et al., 2002; Joo et al., 2017; Konner & Dupont, 2004; Park et al., 2015). The resultant protein forms a Y-shaped homodimer with high binding affinity to VEGFA (the most abundant type of VEGF), VEGFB, and placental growth factor (PlGF) (Konner & Dupont, 2004; Papadopoulos et al., 2012).
(Konner & Dupont, 2004)
The VEGF-Trap fusion protein has been shown to have higher binding affinity, faster association rate, and longer half-life than the VEGF-binding monoclonal antibodies on the market (Papadopoulos et al., 2012). Since the fusion protein seems to work so well in this case, my overarching idea for this project was that maybe fusion proteins with similar structure to VEGF-Trap could be used as decoy receptors in other signaling pathways, too. My goal was to create a Python script that could generate the sequence of a fusion protein similar to VEGF-Trap, given an input of receptor proteins with similar structure to the VEGF receptors.
The VEGF receptors are tyrosine kinases. There are ~60 human receptor tyrosine kinases (RTKs), which can be categorized into ~20 different structural families (Alberts et al., 2015). Other RTK families with extracellular immunoglobulin-like domains besides the VEGF receptors, include the PDGFR, FGFR, CCK, NGFR, EPHR, AXL, TIE, ROR, and MUSK families (Ségaliny et al., 2015). The members of these families are generally all well-annotated on the UniProtKB database, including the locations of the Ig-like domains. Therefore, I created RTKfusion.py script, which utilizes the UniProtKB REST API to programmatically retrieve the sequences of user-specified RTK Ig-domains, and concatenate their sequences to produce a specified fusion protein sequence.
As command line arguments, the RTKfusion.py script takes in a list of gene names/UniProt accession IDs, each with a corresponding Ig domain number. After retrieving the sequences from the UniProtKB database, the script generates the fusion protein sequence by concatenating the signal peptide sequence of the first argument's protein, the sequence of each Ig domain specified as an argument, followed finally by the sequence of the IgG1 Fc domain.
For more information on how to execute the script and the outputs generated, refer to the accompanying README file.
In addition to RTKpython.py and the README.txt file, the supplementary files also include the FASTA output files from two example commands:
Outfile: VGFR1_HUMAN_Ig2_VGFR2_HUMAN_Ig3_fusion.fasta
Outfile: PGFRA_HUMAN_Ig2_PGFRB_HUMAN_Ig5_fusion.fasta
RESULTS
In order to test RTKfusion, I attempted to recreate the sequence for VEGF-Trap using the following command:
python3 RTKfusion.py -g FLT1 Ig2 -g KDR Ig3
This command should have generated the amino acid sequence for the fusion of VEGFR1 signal peptide (encoded by gene FLT1), VEGFR1 Ig-domain 2, VEGFR2 (encoded by gene KDR) Ig-domain 3, and IgG1 Fc domain, from N-terminus to C-terminus (i.e. VEGF-Trap).
The following output was generated:
Execution Successful
VGFR1_HUMAN Signal AA (1, 26), fusion AA 1-26: MVSYWDTGVLLCALLSCLLLTGSSSG
VGFR1_HUMAN Ig2 AA (151, 214), fusion AA 27-90: GRELVIPCRVTSPNITVTLKKFPLDTLIPDGKRIIWDSRKGFIISNATYKEIGLLTCEATVNGH
VGFR2_HUMAN Ig3 AA (224, 320), fusion AA 91-187: YDVVLSPSHGIELSVGEKLVLNCTARTELNVGIDFNWEYPSSKHQHKKLVNRDLKTQSGSEMKKFLSTLTIDGVTRSDQGLYTCAASSGLMTKKNST
IgG1 Fc domain AA (104, 330), fusion AA 189-414: DKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK
>VGFR1_HUMAN_Ig2+VGFR2_HUMAN_Ig2+IgG1_Fc
MVSYWDTGVLLCALLSCLLLTGSSSGGRELVIPCRVTSPNITVTLKKFPLDTLIPDGKRIIWDSRKGFIISNATYKEIGLLTCEATVNGHNKNKTVVIPCLGSISNLNVSLCARYPEKRFVPDGNRISWDSKKGFTIPSYMISYAGMVFCEAKINDEDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK
Compared to the VEGF-Trap sequence I found in a literature search, there were some slight differences:
(Park et al., 2015)
The sequence generated by RTKfusion.py was only 414 amino acids long, compared to the 463 amino acids found in the literature. This is mainly due to the fact that the sequence for VEGFR1-Ig2 used in the literature, extends ~20 amino acids longer on both ends than what is annotated as the beginning/end on UniProtKB.
The script was also tested using several other receptor Ig-domains from various other RTK families (e.g. PDGFRA (P16234) and PDGFRB (P09619)), as well as with variable numbers of gene/Ig-domain pairs, and the script functioned as intended.
As the script retrieves information using HTTP requests, the runtime is dependent on internet service and database server speeds. Additionally, the runtime also varies depending on the number of arguments specified. However, I personally found that with 2 gene arguments, the runtime was typically less than 5 seconds:
real 0m4.011s
user 0m0.266s
sys 0m0.234s
LIMITATIONS AND POTENTIAL IMPROVEMENTS
There are several limitations to the RTKfusion.py script. Firstly, in order to be useful, the user must have extensive prior knowledge of the RTK proteins and where the ligand binding sites are located. Otherwise, the Ig domains chosen may not be involved in ligand binding. In the case of VEGF-Trap, it was experimentally determined that the two receptor Ig domains used could bind the VEGF variants on their own (Wiesmann et al., 2000). However, there is not an annotation on the UniProtKB entries for these receptors indicating that these Ig domains are the main ligand-binding domains, and for many other RTKs, the exact role each Ig domain plays in ligand-binding has not yet been determined. A related potential improvement would be if, instead of having the user input the specific Ig-domain, the script could predict Ig domain primarily responsible for ligand binding, perhaps based on sequence/structural similarity to VEGFR1 Ig2 and VEGFR2 Ig3.
Additionally, the script has only been tested on RTK proteins. It could potentially be expanded to work with other proteins with Ig-like ligand-binding domains. Another limitation is that the script only produces the amino acid sequence. As an improvement, the amino acid sequence could be reversed back to cDNA, which could then be used to construct the expression plasmid vector.
REFERENCES
Alberts, B., Johnson, A., Lewis, J., Morgan, D., Raff, M. C., Roberts, K., Walter, P., Wilson, J. H., & Hunt, T. (2015). Chapter 15: Cell Signaling. In Molecular biology of the cell (6th ed., pp. 813–888). Garland Science.
Hicklin, D. J., & Ellis, L. M. (2005). Role of the vascular endothelial growth factor pathway in tumor growth and angiogenesis. Journal of Clinical Oncology, 23(5), 1011–1027. https://doi.org/10.1200/jco.2005.06.081
Holash, J., Davis, S., Papadopoulos, N., Croll, S. D., Ho, L., Russell, M., Boland, P., Leidich, R., Hylton, D., Burova, E., Ioffe, E., Huang, T., Radziejewski, C., Bailey, K., Fandl, J. P., Daly, T., Wiegand, S. J., Yancopoulos, G. D., & Rudge, J. S. (2002). VEGF-trap: A VEGF blocker with potent antitumor effects. Proceedings of the National Academy of Sciences, 99(17), 11393–11398. https://doi.org/10.1073/pnas.172398299
Joo, K., Park, S. J., Choi, Y., Lee, J. E., Na, Y. M., Hong, H. K., Park, K. H., Kim, H. M., Chung, J.-Y., & Woo, S. J. (2017). Role of the fc region in the vitreous half-life of Anti-VEGF Drugs. Investigative Opthalmology & Visual Science, 58(10), 4261. https://doi.org/10.1167/iovs.17-21813
Konner, J., & Dupont, J. (2004). Use of soluble recombinant decoy receptor vascular endothelial growth factor trap (VEGF trap) to inhibit vascular endothelial growth factor activity. Clinical Colorectal Cancer, 4. https://doi.org/10.3816/ccc.2004.s.013
Papadopoulos, N., Martin, J., Ruan, Q., Rafique, A., Rosconi, M. P., Shi, E., Pyles, E. A., Yancopoulos, G. D., Stahl, N., & Wiegand, S. J. (2012). Binding and neutralization of vascular endothelial growth factor (VEGF) and related ligands by VEGF trap, ranibizumab and bevacizumab. Angiogenesis, 15(2), 171–185. https://doi.org/10.1007/s10456-011-9249-6
Park, S. J., Oh, J., Kim, Y.-K., Park, J. H., Park, J. Y., Hong, H. K., Park, K. H., Lee, J.-E., Kim, H. M., Chung, J. Y., & Woo, S. J. (2015). Intraocular pharmacokinetics of intravitreal vascular endothelial growth factor-trap in a rabbit model. Eye, 29(4), 561–568. https://doi.org/10.1038/eye.2014.329
Ségaliny, A. I., Tellez-Gabriel, M., Heymann, M.-F., & Heymann, D. (2015). Receptor tyrosine kinases: Characterisation, mechanism of action and therapeutic interests for bone cancers. Journal of Bone Oncology, 4(1), 1–12. https://doi.org/10.1016/j.jbo.2015.01.001
Spilsbury, K., Garrett, K. L., Shen, W.-Y., Constable, I. J., & Rakoczy, P. E. (2000). Overexpression of vascular endothelial growth factor (VEGF) in the retinal pigment epithelium leads to the development of choroidal neovascularization. The American Journal of Pathology, 157(1), 135–144. https://doi.org/10.1016/s0002-9440(10)64525-7
Wiesmann, C., Muller, Y. A., & de Vos, A. M. (2000). Ligand-binding sites in ig-like domains of receptor tyrosine kinases. Journal of Molecular Medicine, 78(5), 247–260. https://doi.org/10.1007/s001090000082